About Us
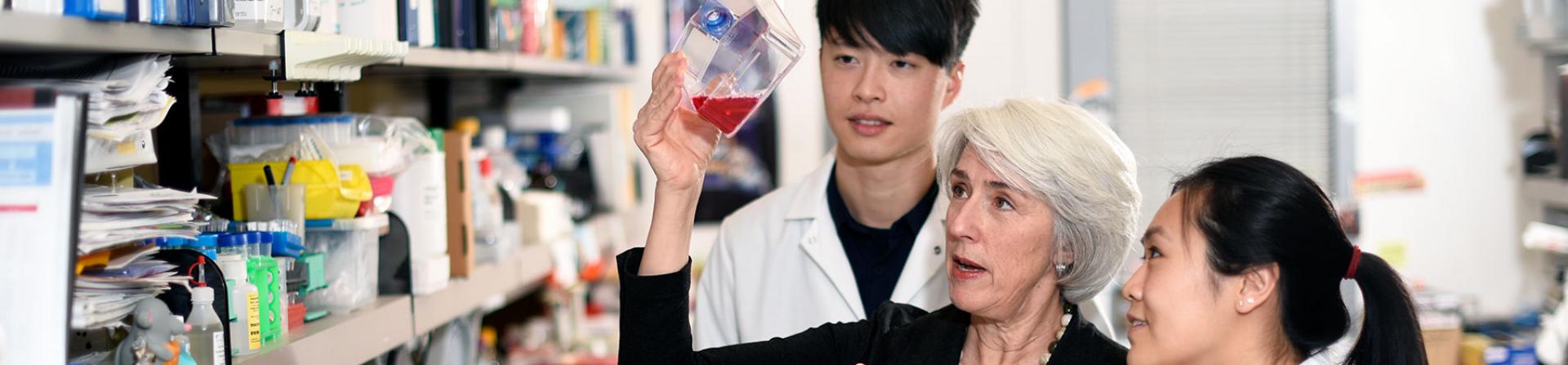
The Department of Molecular and Cellular Biology at Baylor College of Medicine delivers cutting-edge research focused on cancer biology, stem cell biology, developmental biology, and neurobiology. The department ranks among the top ten in the nation in funding from the National Institutes of Health. Our faculty participate in the Dan L Duncan Comprehensive Cancer Center, the Lester and Sue Smith Breast Center, and the Stem Cells and Regenerative Medicine (STaR) Center, among others. The department is home to the Center for Precision Environmental Health. Recruiting junior faculty in both existing and emerging areas of research, such as epigenetics, cancer, and synthetic biology, we attract, train and grow top talent in a collaborative environment with an entrepreneurial spirit.
Message from the Chair
View a message from Department of Molecular and Cellular Biology Chair, Margaret “Peggy” Goodell, Ph.D.
MCB Spotlight
View a series of videos showing what it’s like to start your lab at Baylor College of Medicine within the Texas Medical Center.
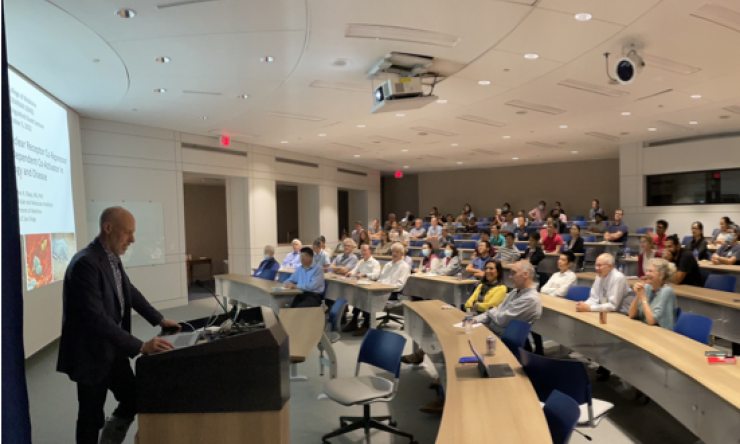
Wednesday Seminar
Join us on Wednesdays to hear from distinguished scientists from around the world, available both in-person and virtually.
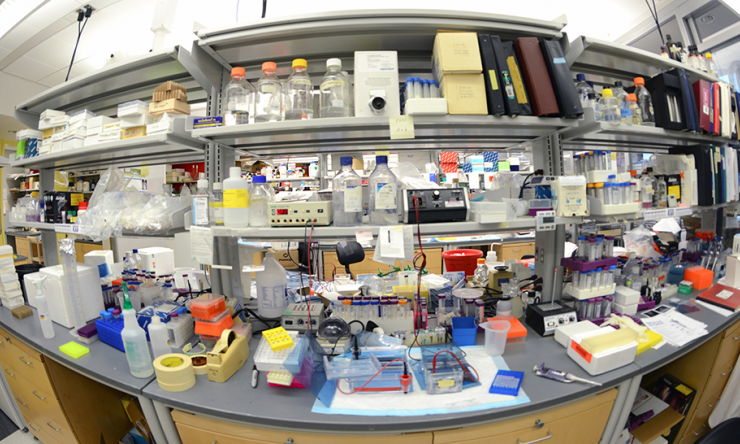
Research
Modern facilities are generously equipped with a full range of instrumentation required for research in cellular, molecular, developmental, and endocrine biology.
Our Faculty
View a listing of the Department of Molecular and Cellular faculty with links to their bios.
Join Our Team
Search for current opportunities offered within the Department of Molecular and Cell Biology.
Diversity, Equity and Inclusion
View information on how the Department of Molecular and Cellular Biology supports our faculty, staff and trainees.