Site Specific Recombinases: Cre & Genetic Reporters
Several site-specific recombinases exist, such as FlpE, Flpo, Phi31c, Cre and now Dre (among others). Our lab's principal interest in these arises from the ability to ablate a gene of interest's function in a temporally and spatially restricted manner (or "conditionally knockout"). For this type of technology, the SSR Cre, and its target sequence loxP, has been the default method of choice for several biologists.
A second reason we are interested in SSRs is because we are curious about cellular differentiation and the contribution of early progenitor cells to the developing embryo and adult. Accordingly, being able to visually identify early progenitors (or distinct populations) of cells and their descendants in a heritable, permanent fashion is highly desirable. This type of experiment is referred to as a genetic lineage tracing. For an excellent review on various lineage tracing strategies, see this paper from Buckingham and Meihlac, 2011.
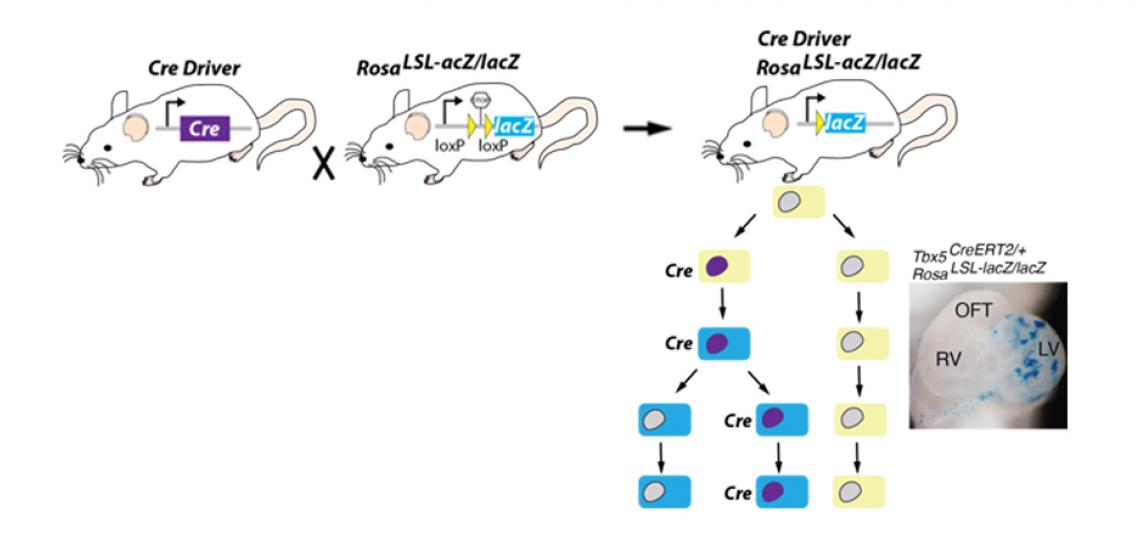
Cre & Genetic Reporters
Several SSR-dependent genetic reporter lines exist, but the lion's share are targeted to the Rosa26 locus and involve a loxP flanked "stop" cassette, or lox-stop-lox (or LSL) element blocking transcription from either the endogenous promoter at the Rosa locus or an artificial CAGGs promoter targeted to the Rosa locus. Thus, when Cre (or another SSR) acts on the sites to recombine out the intervening "stop" cassette, transcription of the reporter (in the image to the right, lacZ) is driven in all cells that express enough Cre molecules to induce recombination.
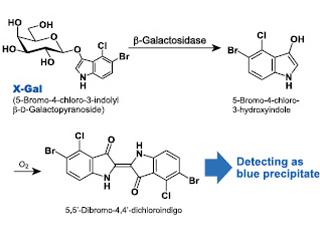
Briefly, for those not familiar, X-gal (5-bromo-4-chloro-3-indoyl-B-d-galactopyranoside) is a colorless, soluble chemical that has a galactose linked to an indole. B-galactosidase (encoded bylacZ) has a high Ka for the galactose part of the substrate and it hydrolyzes X-gal to release the indol, which dimerizes to produce the characteristic blue, insoluble product. Thus, in the presence of X-gal, all cell's that ever expressed enough Cre to recombine the sequence between the loxP sites at the Rosa locus, and all of their genetic descendants, will be labeled blue.







