The Agent
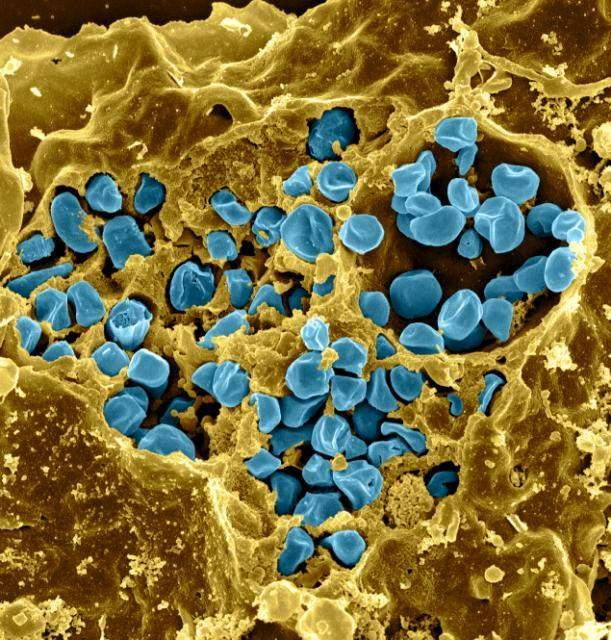
Tularemia is caused by the bacterium Francisella tularensis (F. tularensis), one of the most infectious agents known. Not only does F. tularensis infect a very high number of different species (greater than 250), but an extremely low number of organisms (less than 10) are capable of establishing a potentially lethal infection.
The bacterium is naturally found in small mammals such as rabbits, rodents and hares, and in the insects that feed on these animals (for this reason, the disease is sometimes called rabbit fever or deerfly fever). People can become infected with this bacterium as a result of bites by infected flies, ticks or other insects, handling of infected animal material, eating or drinking contaminated water, or inhaling the bacterium. Tularemia is not known to spread from one person to another.
Cases of infection in the United States are quite rare, with only about 200 cases reported each year, but experts think that some cases of tularemia may not be recognized or reported. Less than two percent of reported cases end in death. The symptoms vary depending on the route of infection and may range from mild illness to severe respiratory illness, including pneumonia and a serious systemic infection. The most dangerous form of the illness is caused by inhalation of the bacterium. Tularemia responds to antibiotics if treated early enough.
There are four subspecies of Francisella tularensis: subspecies tularensis (Type A), subspecies holarctica (Type B), subspecies novicida, and subspecies mediasiatica. Type A is found exclusively in North America and Mexico and causes the most severe form of human disease. Type B is found throughout the Northern Hemisphere and is less pathogenic in humans than Type A.
A vaccine strain was developed several decades ago in the former Soviet Union following repeated passage of a Type B strain. The attenuated, or weakened, live vaccine strain (LVS) offers protection against both Type A and Type B infection. However, because the vaccine strain is not well characterized and the details of how the strain changed to become less virulent in humans are poorly understood, LVS is not licensed for use as a vaccine against tularemia in the United States.
The Problem
Francisella tularensis is classified as a highest risk, Category A potential bioterrorism agent.
Research programs directed toward the development of F. tularensis as a biological weapon previously existed in Japan, the former Soviet Union, and the United States. This research included the creation of strains resistant to existing antibiotics and vaccines, as well as strains harboring virulence genes from other pathogens.
F. tularensis is a formidable threat as a possible bioterrorism agent for the following reasons:
- It can cause an incapacitating, potentially fatal illness.
- Inhalation of the bacterium produces severe respiratory and systemic disease that can result in a mortality rate of 30 percent or more if not treated with antibiotics.
- It is very infectious – as few as 10 organisms can cause disease. It could be easily disseminated in an aerosol form, although the creation of this form of the bacteria requires sophisticated technology.
- It is found widely in nature, so it would not be extremely difficult to obtain the bacterium and grow it under relatively simple laboratory conditions.
- There is no vaccine currently available for general use.
- An outbreak could potentially produce a large burden on the public health system.
The Research
Following the terrorist attacks of 2001, research on Category A pathogens has been made a high priority. Because of the potential risk of F. tularensis as a bioweapon, there is a need to develop safe vaccines, as well as diagnostic tools for this agent.
In addition, researchers need to understand more fully the basic biology of F. tularensis in order to protect humans from a potential threat from F. tularensis. For example, they need to know (1) how it causes disease, (2) which bacterial factors are responsible for its virulence, (3) its growth cycle, and (4) its natural host(s).
Currently, there is little information on many aspects of F. tularensis biology, in part because it is such a severe health threat, so that studies of virulent strains of this organism must be conducted in Biosafety Level 3 (BSL-3) laboratory facilities and are subject to strict regulations and laboratory practices. Furthermore, basic genetic systems to study and alter F. tularensis strains have been developed only recently.
Vaccine Development and Testing
Two vaccines have been studied in humans in the United States – a killed vaccine and a live vaccine strain. Previous studies have shown that the live vaccine strain (LVS) was more effective, but although the LVS has been in use for decades, it has never been licensed for use in humans in the United States due to concerns about its potential to revert to a disease-causing strain and the production methods used to generate the vaccine. Because the supply of this vaccine is limited and aging, a new lot of LVS has been produced using current good manufacturing practices.
Investigators in the Department, including Drs. Hana El Sahly, Wendy Keitel, Robert Atmar, and Shital Patel, have conducted a phase I clinical trial of the new vaccine lot to test the safety, reactogenicity, and immunogenicity of the vaccine in humans.
The goals of their study were to:
- Test the safety and tolerability of increasing doses of LVS among healthy human subjects given by two different routes – under the skin (subcutaneous) or into the skin (scarification).
- Determine the dose of LVS required to infect humans when administered by the two different routes.
- Determine whether the bacterium can be grown from clinical samples collected at various times after administration of LVS at different dosages and routes.
- Analyze immune responses (both humoral and cell-mediated) to LVS.
The results of their study indicated that all doses tested – administered either by the subcutaneous or scarification method – were well tolerated by the healthy human subjects. When compared to subjects receiving a placebo, vaccination by scarification with certain doses of LVS resulted in a significantly higher immune response using a test that measured antibody levels. They also found conditions that elicited significant production of interferon-gamma, a marker for a cell-mediated immune response. Although the study group was small, this trial provided recommendations for further tularemia vaccine evaluation.
To follow up, the investigators expanded their investigation by conducting a phase 2 clinical trial in which 228 participants received the new LVS vaccine by scarification. The safety, reactogenicity, and antibody levels were assessed at multiple time points up to six months, and the results were compared with the existing vaccine that had been in use for decades. This study found that both vaccines were safe and had similar reactogenicity and antibody responses, although the new vaccine was superior for early antibody production.
Genomics of F. tularensis
In addition to vaccine evaluation, researchers in the Department are addressing critical areas of F. tularensis research using an approach called comparative genomics. This work, led by Dr. Joseph Petrosino, involves obtaining the genome sequences of several important strains of F. tularensis. The DNA sequencing is performed using state-of-the-art technology and resources in the Baylor College of Medicine Human Genome Sequencing Center. With this data, the investigators can compare the genomes of different strains to identify F. tularensis virulence factors and potential targets for drug, vaccine and immunodiagnostic development.
The scientists want to determine and compare the genome sequences of multiple F. tularensis strains, including Type A, Type B, and LVS (the weakened version of Type B used as a vaccine). Towards this end, Drs. Petrosino, Nadim Ajami and their colleagues published a high-quality draft genome sequence of a Type B strain in 2015.
Once the comparative genome information is obtained, the researchers can learn how the strains are evolutionarily related, why the Type A strains are more virulent than the Type B strains, why Francisella is so pathogenic, and which mutations weaken LVS and which could weaken pathogenic strains. Using genome comparisons of virulent F. tularensis strains with LVS, they can identify the mutations that attenuate LVS, and then these mutations can be introduced into a virulent Francisella strain and tested for their ability to confer an attenuated phenotype in a mouse tularemia model.
Dr. Petrosino and collaborators are also using functional genomics approaches to identify proteins important for F. tularensis recognition by the host immune system. They are using the genome sequence data to isolate every gene and produce every protein from the Type A and Type B F. tularensis strains. Working with the scientists in the BCM Vaccine Treatment and Evaluation Unit, these proteins will be used in screens to identify the comprehensive set of proteins recognized by the human immune response to F. tularensis.
Results from this work will further characterize the human response to LVS vaccination and lead to the development of novel F. tularensis vaccines and immunodiagnostic reagents. Furthermore, the functional genomic approaches developed in this work with F. tularensis can be adapted to vaccine and diagnostic discovery for other emerging infectious disease agents and potential bioweapons.
For More Information
Learn more about some of the technical terms found on this page by visiting our glossary of terms.