
Join Our Lab!
We are recruiting postdocs and graduate students for new and ongoing projects.
- Graduate Students
- Postdoctoral Associates
Graduate Students:
- Contact Tom Cooper to discuss rotation opportunities.
About the Lab
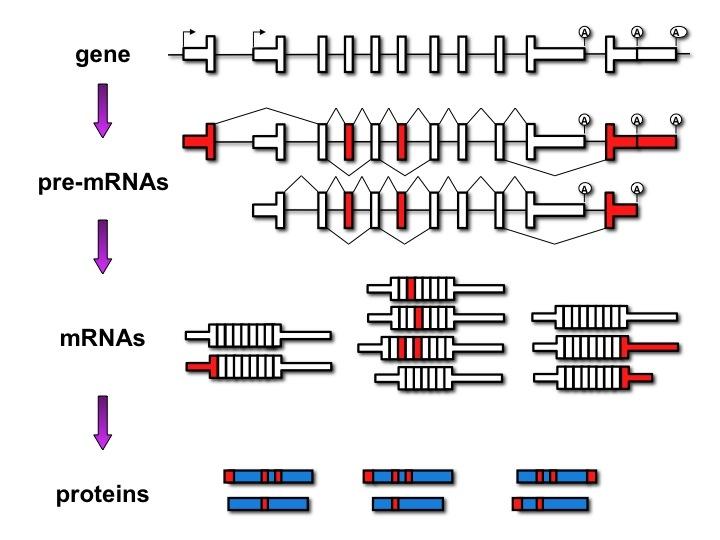
The realization that most human genes generate multiple mRNAs encoding divergent protein isoforms via alternative splicing and polyadenylation has revealed extensive regulation that remains to be explored.
Alternative pathways of RNA processing are regulated in response to signaling cues often coordinating expression of gene networks in response to physiological change including in development and disease.
We are interested in understanding the mechanisms and consequences of this regulation, from how RNA binding proteins and signaling pathways coordinate RNA processing networks to the functional consequences of the different protein isoforms that are expressed in different cell states.
We also investigate the pathogenic mechanisms of myotonic dystrophy, type 1 (DM1), an autosomal dominant neuromuscular disorder affecting multiple tissues including muscle, heart and the central nervous system. The pathogenic mechanism is disruption of developmentally regulated RNA processing, primarily alternative splicing, in which failure to express adult splicing patterns causes primary features of the disease.
The understanding of the molecular mechanisms of DM1 pathogenesis has led to development of several therapeutic approaches some of which are being testing using mouse models established in the lab.
These investigations utilize a combination of cell culture and genetic models including transgenic and knock out mouse lines for RNA binding proteins, CRISPR-derived mouse lines in which specific alternative exons are removed, and DM1 mouse models.
The overlapping areas of investigation in the lab lead to synergistic and collaborative interactions in which knowledge gained in one area fosters progress in the others.
Cooper Lab News
- Rong-Chi’s paper describing the long-term effects of CUGexp RNA expression in a heart model of myotonic dystrophy type 1 was published in J. Clin. Investigation, Insight.
- Larissa has been awarded an NIH K99/R00 award.
- Ashwini received a two-year predoctoral fellowship from the Myotonic Dystrophy Foundation starting January 2026.
- Larissa’s abstract was selected for a lightning round presentation and she won best poster at the 2025 Myotonic Dystrophy Foundation Family Conference.
- Zach Castles from the Genetics and Genomics Graduate Program joined the Cooper lab!
- Rong-Chi Hu, Ph.D. will start her postdoctoral training in the Cooper lab May 2025
- Larissa was awarded the best talk of the postdoc presentations at the Department of Pathology & Immunology Trainee Symposium.
- Larissa’s abstract was selected for a lighting round presentation at the 2025 Myotonic Dystrophy Association Family Conference
- Rong-Chi’s paper describing rescue of the mouse heart model for myotonic dystrophy by muscleblind expression was published in the J. Clinical Investigation, 2025
- Lathan defended his thesis and graduated 2025
- Jesus Frias, Ph.D. joined our lab as a new postdoctoral fellow, 2024
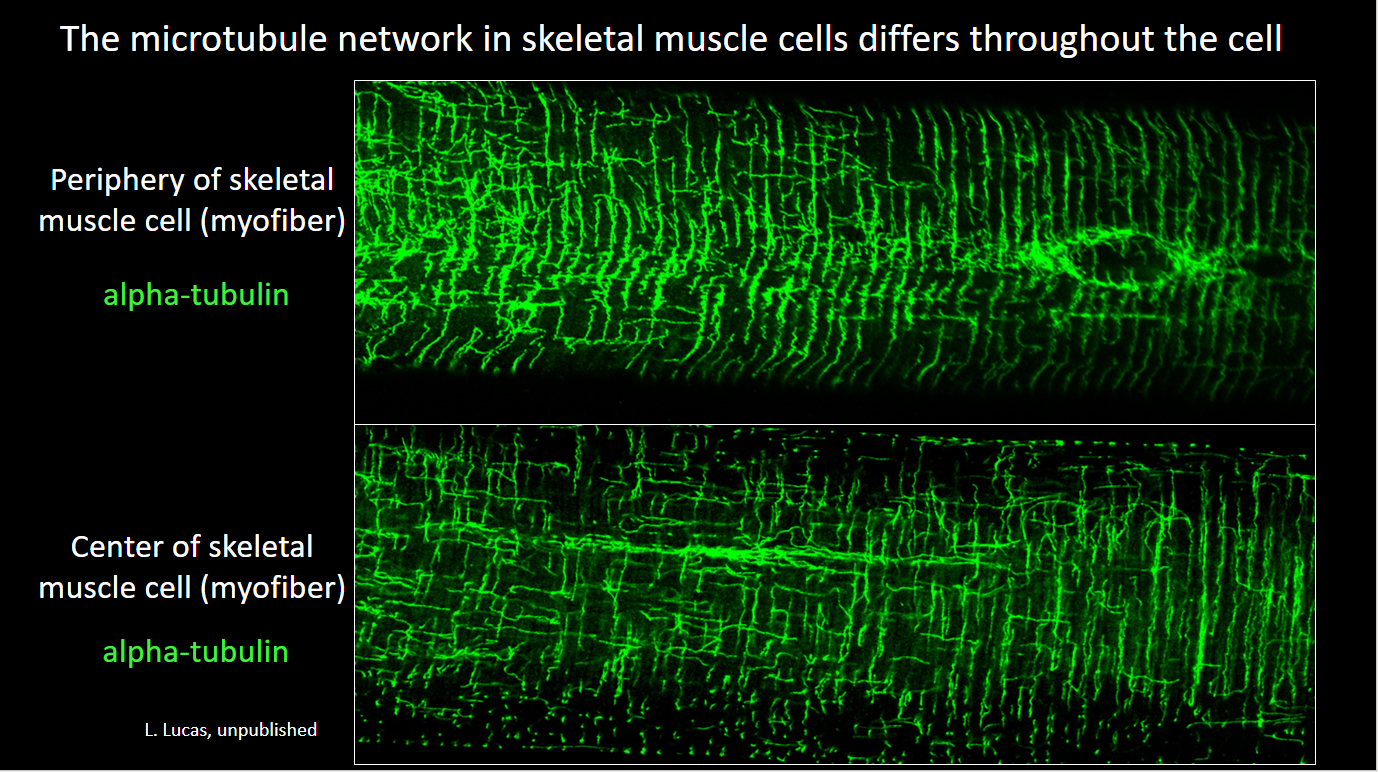
- Lathan’s paper on the role of the striated muscle splice variant of Map4 in skeletal muscle microtubule structure was published in iScience, 2024
- Ashwini joined our lab as a new Ph.D. student, 2024
- Ashwini is an awardee of the Trentin Award for academic achievement 2024
- Sara’s paper that identifies HSP90 as a modifier of RNA foci in myotonic dystrophy type was published in Mol. Cell Biol. 2024
- Sara defended her thesis and graduated 2024
Reagents
The following plasmids are available through Addgene.
FRE5 (FLAGNLSREDGFP5)
FRE5Cf (FLAGNLSREDGFP5Claf)
RG6
RHCglo
RTB300
R300TA
DT960
DT480
DT240
DT40
DT12A
DT0
pBItetDT960GFP
pBItetDT12nGFP
FLYLQ (Flag-CELF1)
pBItetDMPKSGFP
pBItetDT240GFP
pBItetDT480GFP
FlagETR3 (CELF2)
FlagMBNL1-41
FlagMBNL3
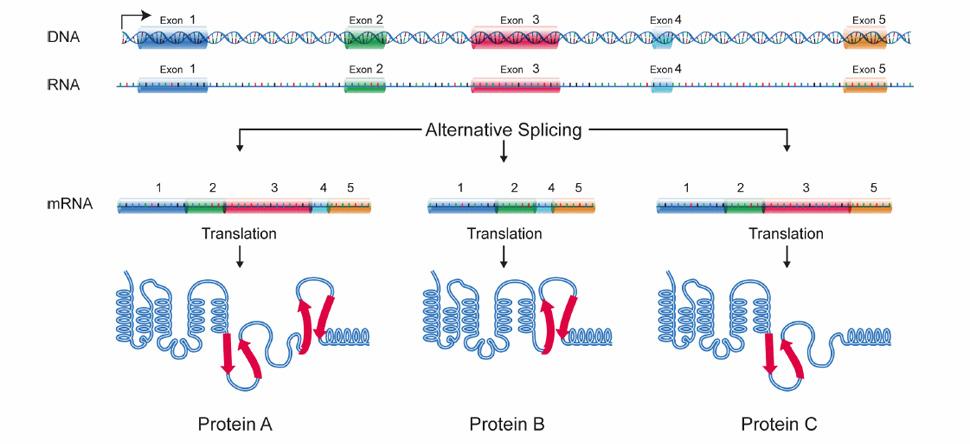
Alternative splicing is crucial to muscle mass maintenance
Alternative splicing allows cells to make many different proteins with a limited number of genes.
There is more going on in DM1 than just alternative splicing
In his laboratory at Baylor College of Medicine, Dr. Thomas A. Cooper is leading the way to better understand myotonic dystrophy type 1 (DM1), a rare but devastating condition.

What causes myotonic dystrophy, type1
The lab of Dr. Thomas Cooper discovered that the CUGexp RNA disrupts the regulation of RNA splicing that is important for proper functions of a large number of genes.
Image of the Month: From the Labs blog