Image of the Month: October 2022
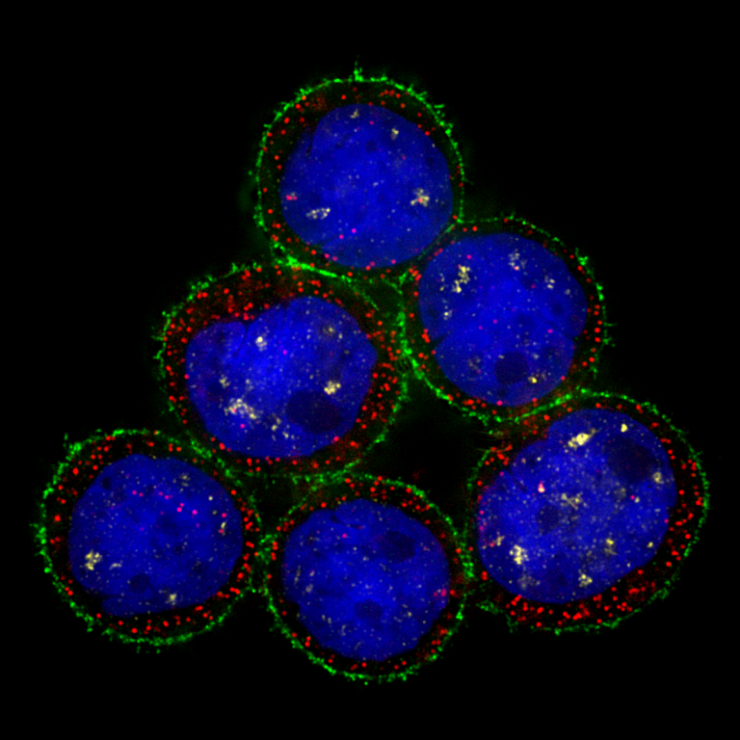
High resolution deconvolution microscopy-based image of MDA-MB-453 human breast cancer cells simultaneously labeled by immunofluorescence (Her-2, Green), DNA (DAPI, Blue, nucleus) and mRNA FISH (Malat1, Yellow; GAPDH, Red).
Each month the Baylor College of Medicine blog, From the Labs, features an image from our labs and cores. For the month of October 2022, the Integrated Microscopy Core's image was published in the blog. This image also represents BCM's research efforts durning Breast Cancer Awareness Month by featuring human breast cancer cells.
About the Core
The Integrated Microscopy Core is an institutional core facility supported by the Advanced Technology Cores at Baylor College of Medicine, the NCI Comprehensive Dan L Duncan Cancer Center, the TMC Digestive Disease Center and Cancer Prevention and Research Institute of Texas.
The core is designed to provide high-quality microscopy and analytics services through availability of state-of-the-art imaging instrumentation, image analysis, one-on-one training by expert staff, plus free consultation for experimental setup, data acquisition and analysis.
The IMC covers most of the investigators’ needs in the areas of light microscopy with particular emphasis on high throughput/high content phenotypic screening (assay development/validation), live imaging, multi-dimensional spatial analysis in tissue and cell culture samples, and 3D culture models imaging.
Equipment and Services
- Equipment available
- Free consultation in experimental design and data interpretation (by appointment only)
- Tech assist in experiment setup, execution and imaging (limited)
- Assisted Use on all instruments
- Laminar flow hood and CO2 incubator for limited-time cell maintenance and live experiments (virus or bacteria require proper approval)
- High Throughput Microscopy experiments
- Image Analysis
Publication Acknowledgement for Cores
Core name: Integrated Microscopy Core
Personnel: Michael A. Mancini, Ph.D., Academic Director; Michael Bolt, Ph.D., Core Director; Elina Mosa
Grants supporting the core: The Integrated Microscopy Core is supported by the Center for Advanced Microscopy and Image Informatics (CAMII) with funding from NIH (DK56338, CA125123, ES030285), and CPRIT (RP150578, RP170719).
Publications
Deater M, Tamhankar M, Lloyd RE. TDRD3 is an antiviral restriction factor that promotes IFN signaling with G3BP1. PLoS Pathog. 2022 Jan 27;18(1):e1010249. doi: 10.1371/journal.ppat.1010249. PMID: 35085371; PMCID: PMC8824378.
Doerfler AM, Han J, Jarrett KE, Tang L, Jain A, Saltzman A, De Giorgi M, Chuecos M, Hurley AE, Li A, Morand P, Ayala C, Goodlett DR, Malovannaya A, Martin JF, de Aguiar Vallim TQ, Shroyer N, Lagor WR. Intestinal Deletion of 3-Hydroxy-3-Methylglutaryl-Coenzyme A Reductase Promotes Expansion of the Resident Stem Cell Compartment. Arterioscler Thromb Vasc Biol. 2022 Apr;42(4):381-394. doi: 10.1161/ATVBAHA.122.317320. Epub 2022 Feb 17. PMID: 35172604; PMCID: PMC8957608.
Junco JJ, Zorman B, Gant VU Jr, Muñoz J, Lacorazza HD, Sumazin P, Rabin KR. CRLF2 overexpression results in reduced B-cell differentiation and upregulated E2F signaling in the Dp16 mouse model of Down syndrome. Exp Hematol. 2022 Mar 17:S0301-472X(22)00127-8. doi: 10.1016/j.exphem.2022.03.005. Epub ahead of print. PMID: 35306048.
Li Q, Holliday M, Pan JS, Tan L, Li J, Sheikh-Hamad D. Interactions between leucines within the signal peptides of megalin and stanniocalcin-1 are crucial for regulation of mitochondrial metabolism. Lab Invest. 2022 May;102(5):534-544. doi: 10.1038/s41374-022-00729-3. Epub 2022 Jan 19. PMID: 35046485.
Moree SE, Maneix L, Iakova P, Stossi F, Sahin E, Catic A. Imaging-Based Screening of Deubiquitinating Proteases Identifies Otubain-1 as a Stabilizer of c-MYC. Cancers (Basel). 2022 Feb 4;14(3):806. doi: 10.3390/cancers14030806. PMID: 35159073; PMCID: PMC8833929.
Rajan A, Piedra FA, Aideyan L, McBride T, Robertson M, Johnson HL, Aloisio GM, Henke D, Coarfa C, Stossi F, Menon VK, Doddapaneni H, Muzny DM, Javornik Cregeen SJ, Hoffman KL, Petrosino J, Gibbs RA, Avadhanula V, Piedra PA. Multiple Respiratory Syncytial Virus (RSV) Strains Infecting HEp-2 and A549 Cells Reveal Cell Line-Dependent Differences in Resistance to RSV Infection. J Virol. 2022 Apr 13;96(7):e0190421. doi: 10.1128/jvi.01904-21. Epub 2022 Mar 14. PMID: 35285685; PMCID: PMC9006923.
Rajan A, Weaver AM, Aloisio GM, Jelinski J, Johnson HL, Venable SF, McBride T, Aideyan L, Piedra FA, Ye X, Melicoff-Portillo E, Yerramilli MRK, Zeng XL, Mancini MA, Stossi F, Maresso AW, Kotkar SA, Estes MK, Blutt S, Avadhanula V, Piedra PA. The Human Nose Organoid Respiratory Virus Model: an Ex Vivo Human Challenge Model To Study Respiratory Syncytial Virus (RSV) and Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Pathogenesis and Evaluate Therapeutics. mBio. 2022 Feb 15;13(1):e0351121. doi: 10.1128/mbio.03511-21. Epub ahead of print. PMID: 35164569; PMCID: PMC8844923.
Robichaux MA, Nguyen V, Chan F, Kailasam L, He F, Wilson JH, Wensel TG. Subcellular localization of mutant P23H rhodopsin in an RFP fusion knock-in mouse model of retinitis pigmentosa. Dis Model Mech. 2022 May 1;15(5):dmm049336. doi: 10.1242/dmm.049336. Epub 2022 May 6. PMID: 35275162.
Singh S, Lee N, Pedroza DA, Bado IL, Hamor C, Zhang L, Aguirre S, Hu J, Shen Y, Xu Y, Gao Y, Zhao N, Chen SH, Wan YW, Liu Z, Chang JT, Hollern D, Perou CM, Zhang XHF, Rosen JM. Chemotherapy coupled to macrophage inhibition induces T-cell and B-cell infiltration and durable regression in triple-negative breast cancer. Cancer Res. 2022 Apr 20:canres.3714.2021. doi: 10.1158/0008-5472.CAN-21-3714. Epub ahead of print. PMID: 35442423.
Stossi F, Singh PK, Mistry RM, Johnson HL, Dandekar RD, Mancini MG, Szafran AT, Rao AU, Mancini MA. Quality Control for Single Cell Imaging Analytics Using Endocrine Disruptor-Induced Changes in Estrogen Receptor Expression. Environ Health Perspect. 2022 Feb;130(2):27008. doi: 10.1289/EHP9297. Epub 2022 Feb 15. PMID: 35167326; PMCID: PMC8846386.
Bolt MJ, Singh P, Obkirchner CE, Powell RT, Mancini MG, Szafran AT, Stossi F, Mancini MA. Endocrine disrupting chemicals differentially alter intranuclear dynamics and transcriptional activation of estrogen receptor-α. iScience. 2021 Oct 7;24(11):103227. doi: 10.1016/j.isci.2021.103227. PMID: 34712924; PMCID: PMC8529556.
Zhao N, Powell RT, Yuan X, Bae G, Roarty KP, Stossi F, Strempfl M, Toneff MJ, Johnson HL, Mani SA, Jones P, Stephan CC, Rosen JM. Morphological screening of mesenchymal mammary tumor organoids to identify drugs that reverse epithelial-mesenchymal transition. Nat Commun. 2021 Jul 12;12(1):4262. doi: 10.1038/s41467-021-24545-3. PMID: 34253738; PMCID: PMC8275587.
Bomidi C, Robertson M, Coarfa C, Estes MK, Blutt SE. Single-cell sequencing of rotavirus-infected intestinal epithelium reveals cell-type specific epithelial repair and tuft cell infection. Proc Natl Acad Sci U S A. 2021 Nov 9;118(45):e2112814118. doi: 10.1073/pnas.2112814118. PMID: 34732579; PMCID: PMC8609316.
Cox AR, Chernis N, Kim KH, Masschelin PM, Saha PK, Briley SM, Sharp R, Li X, Felix JB, Sun Z, Moore DD, Pangas SA, Hartig SM. Ube2i deletion in adipocytes causes lipoatrophy in mice. Mol Metab. 2021 Jun;48:101221. doi: 10.1016/j.molmet.2021.101221. Epub 2021 Mar 24. PMID: 33771728; PMCID: PMC8080079.
Hebert KA, Jaramillo S, Yu W, Wang M, Veeramachaneni R, Sandulache VC, Sikora AG, Bonnen MD, Annapragada AV, Corry D, Kheradmand F, Pandita RK, Ludwig MS, Pandita TK, Huang S, Coarfa C, Grimm SL, Perera D, Miles G, Ghebre YT. Esomeprazole enhances the effect of ionizing radiation to improve tumor control. Oncotarget. 2021 Jul 6;12(14):1339-1353. doi: 10.18632/oncotarget.28008. PMID: 34262645; PMCID: PMC8274720.
Lee JH, Wang R, Xiong F, Krakowiak J, Liao Z, Nguyen PT, Moroz-Omori EV, Shao J, Zhu X, Bolt MJ, Wu H, Singh PK, Bi M, Shi CJ, Jamal N, Li G, Mistry R, Jung SY, Tsai KL, Ferreon JC, Stossi F, Caflisch A, Liu Z, Mancini MA, Li W. Enhancer RNA m6A methylation facilitates transcriptional condensate formation and gene activation. Mol Cell. 2021 Aug 19;81(16):3368-3385.e9. doi: 10.1016/j.molcel.2021.07.024. Epub 2021 Aug 9. PMID: 34375583; PMCID: PMC8383322.
Marom R, Burrage LC, Venditti R, Clément A, Blanco-Sánchez B, Jain M, Scott DA, Rosenfeld JA, Sutton VR, Shinawi M, Mirzaa G, DeVile C, Roberts R, Calder AD, Allgrove J, Grafe I, Lanza DG, Li X, Joeng KS, Lee YC, Song IW, Sliepka JM, Batkovskyte D, Washington M, Dawson BC, Jin Z, Jiang MM, Chen S, Chen Y, Tran AA, Emrick LT, Murdock DR, Hanchard NA, Zapata GE, Mehta NR, Weis MA, Scott AA, Tremp BA, Phillips JB, Wegner J, Taylor-Miller T, Gibbs RA, Muzny DM, Jhangiani SN, Hicks J, Stottmann RW, Dickinson ME, Seavitt JR, Heaney JD, Eyre DR; Undiagnosed Diseases Network, Westerfield M, De Matteis MA, Lee B. COPB2 loss of function causes a coatopathy with osteoporosis and developmental delay. Am J Hum Genet. 2021 Sep 2;108(9):1710-1724. doi: 10.1016/j.ajhg.2021.08.002. Epub 2021 Aug 26. PMID: 34450031; PMCID: PMC8456174.
Moraes RM, Elefteriou F, Anbinder AL. Response of the periodontal tissues to β-adrenergic stimulation. Life Sci. 2021 Sep 15;281:119776. doi: 10.1016/j.lfs.2021.119776. Epub 2021 Jun 27. PMID: 34186048; PMCID: PMC8319150.
Potter VL, Moye AR, Robichaux MA, Wensel TG. Super-resolution microscopy reveals photoreceptor-specific subciliary location and function of ciliopathy-associated protein CEP290. JCI Insight. 2021 Oct 22;6(20):e145256. doi: 10.1172/jci.insight.145256. PMID: 34520396; PMCID: PMC8564900.
Singh VP, Pinnamaneni JP, Pugazenthi A, Sanagasetti D, Mathison M, Martin JF, Yang J, Rosengart TK. Hippo Pathway Effector Tead1 Induces Cardiac Fibroblast to Cardiomyocyte Reprogramming. J Am Heart Assoc. 2021 Dec 21;10(24):e022659. doi: 10.1161/JAHA.121.022659. Epub 2021 Dec 10. PMID: 34889103.
Cox DC, Guan X, Xia Z, Cooper TA. Increased nuclear but not cytoplasmic activities of CELF1 protein leads to muscle wasting. Hum Mol Genet. 2020 Jun 27;29(10):1729-1744. doi: 10.1093/hmg/ddaa095. PMID: 32412585; PMCID: PMC7322576.
Criglar JM, Crawford SE, Zhao B, Smith HG, Stossi F, Estes MK. A Genetically Engineered Rotavirus NSP2 Phosphorylation Mutant Impaired in Viroplasm Formation and Replication Shows an Early Interaction between vNSP2 and Cellular Lipid Droplets. J Virol. 2020 Jul 16;94(15):e00972-20. doi: 10.1128/JVI.00972-20. PMID: 32461314; PMCID: PMC7375380.
Lin SC, Qu L, Ettayebi K, Crawford SE, Blutt SE, Robertson MJ, Zeng XL, Tenge VR, Ayyar BV, Karandikar UC, Yu X, Coarfa C, Atmar RL, Ramani S, Estes MK. Human norovirus exhibits strain-specific sensitivity to host interferon pathways in human intestinal enteroids. Proc Natl Acad Sci U S A. 2020 Sep 22;117(38):23782-23793. doi: 10.1073/pnas.2010834117. Epub 2020 Sep 9. PMID: 32907944; PMCID: PMC7519316.
Mullany LK, Rohira AD, Leach JP, Kim JH, Monroe TO, Ortiz AR, Stork B, Gaber MW, Sarkar P, Sikora AG, Rosengart TK, York B, Song Y, Dacso CC, Lonard DM, Martin JF, O'Malley BW. A steroid receptor coactivator stimulator (MCB-613) attenuates adverse remodeling after myocardial infarction. Proc Natl Acad Sci U S A. 2020 Dec 8;117(49):31353-31364. doi: 10.1073/pnas.2011614117. Epub 2020 Nov 23. PMID: 33229578.
Muscarella AM, Dai W, Mitchell PG, Zhang W, Wang H, Jia L, Stossi F, Mancini MA, Chiu W, Zhang XH. Unique cellular protrusions mediate breast cancer cell migration by tethering to osteogenic cells. NPJ Breast Cancer. 2020 Sep 7;6:42. doi: 10.1038/s41523-020-00183-8. PMID: 32964116; PMCID: PMC7477119.
Nguyen TM, Kabotyanski EB, Reineke LC, Shao J, Xiong F, Lee JH, Dubrulle J, Johnson H, Stossi F, Tsoi PS, Choi KJ, Ellis AG, Zhao N, Cao J, Adewunmi O, Ferreon JC, Ferreon ACM, Neilson JR, Mancini MA, Chen X, Kim J, Ma L, Li W, Rosen JM. The SINEB1 element in the long non-coding RNA Malat1 is necessary for TDP-43 proteostasis. Nucleic Acids Res. 2020 Mar 18;48(5):2621-2642. doi: 10.1093/nar/gkz1176. PMID: 31863590; PMCID: PMC7049706.
Singh VP, Pinnamaneni JP, Pugazenthi A, Sanagasetti D, Mathison M, Wang K, Yang J, Rosengart TK. Enhanced Generation of Induced Cardiomyocytes Using a Small-Molecule Cocktail to Overcome Barriers to Cardiac Cellular Reprogramming. J Am Heart Assoc. 2020 Jun 16;9(12):e015686. doi: 10.1161/JAHA.119.015686. Epub 2020 Jun 5. PMID: 32500803; PMCID: PMC7429035.
Stossi F, Dandekar RD, Mancini MG, Gu G, Fuqua SAW, Nardone A, De Angelis C, Fu X, Schiff R, Bedford MT, Xu W, Johansson HE, Stephan CC, Mancini MA. Estrogen-induced transcription at individual alleles is independent of receptor level and active conformation but can be modulated by coactivators activity. Nucleic Acids Res. 2020 Feb 28;48(4):1800-1810. doi: 10.1093/nar/gkz1172. PMID: 31930333; PMCID: PMC7039002.
Yu W, Chen Y, Putluri N, Coarfa C, Robertson MJ, Putluri V, Stossi F, Dubrulle J, Mancini MA, Pang JC, Nguyen T, Baluya D, Myers JN, Lai SY, Sandulache VC. Acquisition of Cisplatin Resistance Shifts Head and Neck Squamous Cell Carcinoma Metabolism toward Neutralization of Oxidative Stress. Cancers (Basel). 2020 Jun 24;12(6):1670. doi: 10.3390/cancers12061670. PMID: 32599707; PMCID: PMC7352569.
Zheng ZY, Anurag M, Lei JT, Cao J, Singh P, Peng J, Kennedy H, Nguyen NC, Chen Y, Lavere P, Li J, Du XH, Cakar B, Song W, Kim BJ, Shi J, Seker S, Chan DW, Zhao GQ, Chen X, Banks KC, Lanman RB, Shafaee MN, Zhang XH, Vasaikar S, Zhang B, Hilsenbeck SG, Li W, Foulds CE, Ellis MJ, Chang EC. Neurofibromin Is an Estrogen Receptor-α Transcriptional Co-repressor in Breast Cancer. Cancer Cell. 2020 Mar 16;37(3):387-402.e7. doi: 10.1016/j.ccell.2020.02.003. Epub 2020 Mar 5. PMID: 32142667; PMCID: PMC7286719