About the Center
The Dan L Duncan Comprehensive Cancer Center is a comprehensive cancer center designated by the National Cancer Institute. The DLDCCC combines all our cancer research and treatment capabilities under one umbrella. The comprehensive designation was achieved through our significant accomplishments in career enhancement (education of future doctors and scientists) and community outreach to reduce health disparities and improve survival from cancer, especially in underserved populations.

Research Programs
Learn more about the research programs and the innovative clinical research being conducted in the Dan L Duncan Comprehensive Cancer Center.
Training Programs
We offer various cancer-relevant T32 training awards, training grants funded by the Cancer Prevention and Research Institute of Texas (CPRIT) and competitive training programs.

Letter from Our Director
There is nothing simple about establishing a legacy of integrity, excellence and belonging, yet this is what the Dan L Duncan Comprehensive Cancer Center has achieved for all Houstonians suffering from cancer.
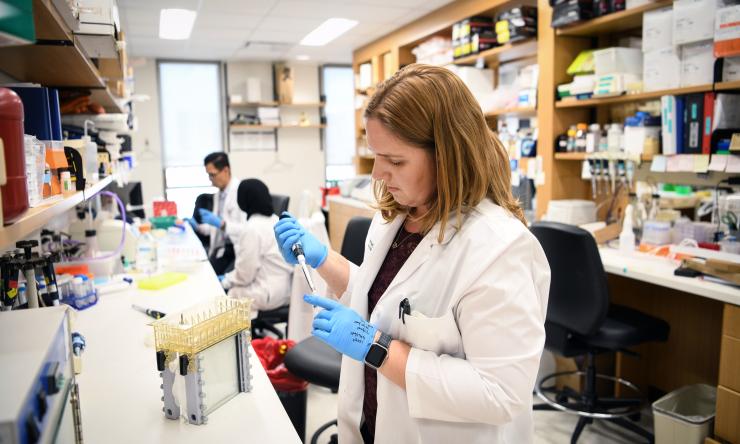

Shared Resources
Check out our core services available to all Duncan Cancer Center members.

Membership
The Duncan Cancer Center has over 450 members in both research and clinical areas.


