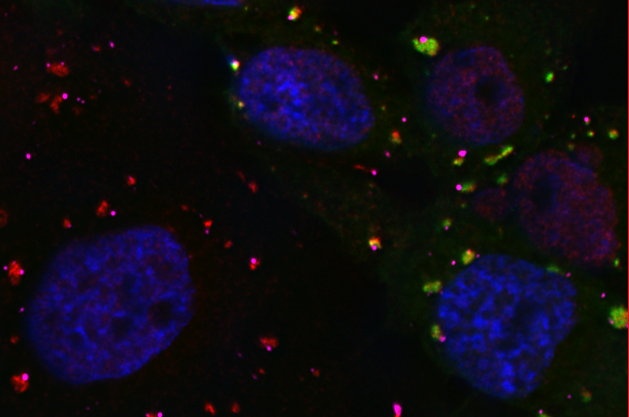
We are interested in mechanisms that regulate RNA Stress Granule (SG) and Processing Body (P-body) formation in cells. These two types of RNA granules aggregate translationally-silenced mRNAs and function as an extension of translation regulation and are key structures that regulate the mRNA cycle. Translational silencing by microRNAs is thought to regulate expression of about 50 percent of human genes, and function by through the SG and P-body pathways.
We found that poliovirus 3Cpro cleaves a protein called G3BP that is critical for nucleating SG formation. We have also shown that P-bodies are disrupted in virus-infected cells and are currently focusing on virus-induced degradation of three key components of P-bodies, Xrn1, Dcp1a and PAN3, all of which regulate mRNA decay in cells. Though we use a virus model system, much of our work also centers on regulation mechanisms in uninfected cells to determine how G3BP actually regulates stress granule formation, and what the mechanisms of RNA granule nucleation are.
We are collaborating with Dr. Mark Bedford at UT MD Anderson to determine the function of several methylation sites on G3BP and the role they play in stress granule assembly and innate immune signaling by G3BP.








