Chonghui Cheng Lab
Cheng lab members celebrating their COVID vaccinations. View a listing of our members.
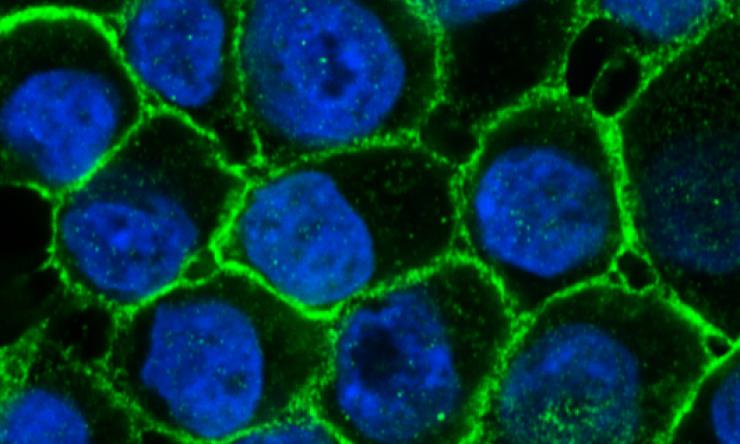
Image of the Month
Human breast epithelial cells are organized as a cobblestone layer revealed here by e-cadherin (epithelial-cadherin, green), a cell-adhesion protein located on the cell surface. Cell nuclei are highlighted in blue. Image courtesy of Nature Communications.

Pink for Breast Cancer
Members of the Cheng Lab showed their support for Breast Cancer Awareness month in October 2019 by donning PINK!

Lab Meeting
Cheng Lab members convene to discuss their research projects.

Lab Outing
Several members of the Cheng lab enjoy the fresh air on an outing the Hermann Park near the Baylor College of Medicine campus.
About the Lab
In the Cheng lab, we strive to understand the fundamental questions of how RNA regulation controls cellular processes in normal biology and in the context of cancer. Working at the interface of RNA splicing and breast cancer biology, our current focus is on regulation of breast cancer metastasis driven by alternative splicing. We use molecular biology, genomics, and bioinformatics approaches in conjunction with genetic models and patient samples to discover rules and networks that regulate metastasis and associated processes. We work closely with physician scientists and aim to apply our findings from basic research to the development of prognostic markers and therapeutics for the treatment of breast cancer.
The developmental program Epithelial-Mesenchymal Transition (EMT) is frequently re-activated in metastatic and recurrent tumors. Our work provided a conceptual understanding depicting a causal role for RNA alternative splicing in EMT and breast cancer recurrence. We found that splice isoform switching of the CD44 gene must take place in order for cells to undergo EMT. We also discovered a novel splicing-mediated pathway that drives cancer metastasis. We demonstrated that the RNA binding protein hnRNPM reprograms alternative splicing including CD44 and promotes a breast cancer metastatic phenotype. By competitive binding on cis-regulatory RNA elements, hnRNPM activates a mesenchymal splicing program in a cell-type restricted manner, emphasizing a tightly regulated splicing program during tumor metastasis. We are combining patient data biocomputing analysis with cell-based and animal experiments to determine the networks of RNA regulation that governs the phenotype of breast cancer metastasis.
In collaboration with nano-technology engineers, we developed the “NanoFlare” method that enables the detection and isolation of live circulating tumor cells (CTC), establishing a platform to study splicing-mediated cancer cell plasticity and phenotypes in patient-derived samples. We are continuing on this collaboration to develop novel tools for the prognosis and diagnosis of breast cancer.
We have been intrigued by the fact that nearly all human genes are detected to undergo alternative splicing, vastly expanding the human proteomes. Therapeutic resistance of promising anti-tumor drugs, such as the anti-HER2 antibody Trastuzumab and the B-RAF(V600E) inhibitor Vemurafenib, is now known to be caused by aberrantly spliced HER2 and B-RAF. Despite these important observations, alternative splicing in cancer has remained largely an untargeted territory. We are actively looking for dedicated research fellows to join us to understand the contribution of RNA regulation in breast cancer metastasis and to apply it to clinical settings.
Cheng Lab
BCM-Alkek Graduate School
Room N1110.02
MS: BCM600
Houston, TX 77030
Phone: 713-798-1056
Email: Chonghui.Cheng@bcm.edu
