Until recently, our understanding of the genetic causes for human diseases were largely limited to the study of extreme phenotypes produced by Mendelian disorders. Advances in genomic technologies and statistical methods have converged to allow the discovery of susceptibility loci for complex genetic traits, and the field now confronts the enormous challenge to confirm the responsible genes and define their functions in the biology of health and disease. In addition to the potential for fundamental biological insights, functional dissection of susceptibility genes will be essential to realize the full promise of human genetics for clinical applications. In the coming years, with the completion of the largest possible genetic meta-analyses along with the advent of whole-genome sequencing, our knowledge of common and rare susceptibility variants will rapidly expand, implicating vast genetic networks including hundreds of loci in neurologic diseases. Innovative strategies are therefore urgently needed to accelerate functional studies.
Background and Significance: The power of disease endophenotypes
Gene discovery in Alzheimer's disease (AD) and Parkinson's disease (PD) has largely relied on the case-control design based on clinical diagnoses; however, this approach is potentially limited by etiologic heterogeneity amongst patients and the presence of substantial but sub-clinical pathology in controls. A complementary approach directly leverages the underlying disease pathology as an intermediate trait, or endophenotype. The clinical manifestation of neurodegenerative disease is the culmination of a multi-tiered pathogenic cascade that evolves over decades—understanding how genetic variants impact this causal chain is essential. Substantial evidence supports a model in which many genetic variants promote the development of neuropathology, subsequently leading to the clinical manifestations of disease.
Pathological Mutations in Alzheimers Disease
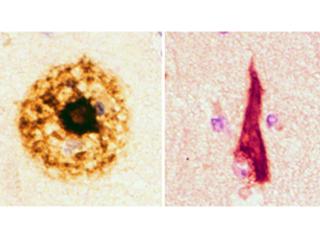
Most notably, rare pathogenic mutations have been identified in the amyloid precursor protein, microtubule associate protein tau, and alpha-synuclein genes, encoding the primary constituents of the defining disease pathologies (amyloid plaques and Tau neurofibrillary tangles in AD, and Lewy bodies in PD, respectively). We have participated in several studies that further illustrate the power of endophenotypes for functional genetic dissection of AD (Shulman et al. 2010; De Jager, Shulman et al. 2012). We have also contributed to genome-wide association studies (GWAS) seeking novel susceptibility loci for AD neuropathologic traits, including amyloid neuritic plaques and Tau neurofibrillary tangles. Our recent work finds that the association of several genes with AD susceptibility, including CR1, CD2AP, and ABCA7, is likely mediated by an impact on underlying amyloid pathology; and further, our genome scan implicates common variation in the amyloid precursor protein (APP) gene (Shulman et al. in press).
Defining the Pathology of Parkinson's Disease
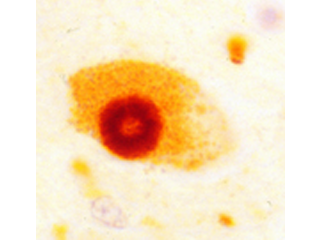
Although two percent of the population over age 65 are clinically diagnosed with PD, the defining pathology of disease (alpha-synuclein Lewy bodies) is discovered in 20 percent of brains from population-based autopsy studies. We have participated in studies to better understand the clinical impact of this pathology, and its potential as an endophenotype for genetic analyses. In a study of 744 autopsies from older adults without a clinical diagnosis of PD, nearly a third showed evidence of post-mortem Lewy bodies and/or substantia nigra neurodegeneration, the defining pathologies of PD, and these changes were associated with parkinsonian motor signs proximate to death (Buchman et al. 2011). Therefore, PD-related pathology appears to be responsible for a much greater burden of age-related motor disability than currently realized. Discovery of the underlying genetic factors may allow us to identify susceptible individuals and ultimately intervene with effective therapies.
Current Projects
We continue to leverage endophenotypes for gene discovery and functional genomic dissection in neurodegenerative diseases, collaborating closely with Dr. David Bennett (Rush University Medical Center) and Dr. Philip De Jager (Brigham and Women's Hospital, Harvard Medical School, and the Broad Institute). Our goal is to perform joint meta-analyses in a combined cohort of more than 2,000 prospectively acquired brain autopsies. These efforts will substantially enhance the power of our genetic analyses of AD pathology traits. In addition, we are applying the endophenotype strategy to the functional investigation and discovery of PD susceptibility genes. First, we are investigating the large number of emerging PD loci for associations with endophenotypes, including alpha-synuclein Lewy bodies, substantia nigra neuronal loss, and the development of parkinsonian motor signs. Second, we are performing a genome-wide scan in our collaborative autopsy cohort to identify susceptibility loci for Lewy body pathology. Discovered variants will be interrogated for their impact along the causal chain of disease, evaluating associations for nigral degeneration and parkinsonism, and we will also determine if these loci impact clinical PD.
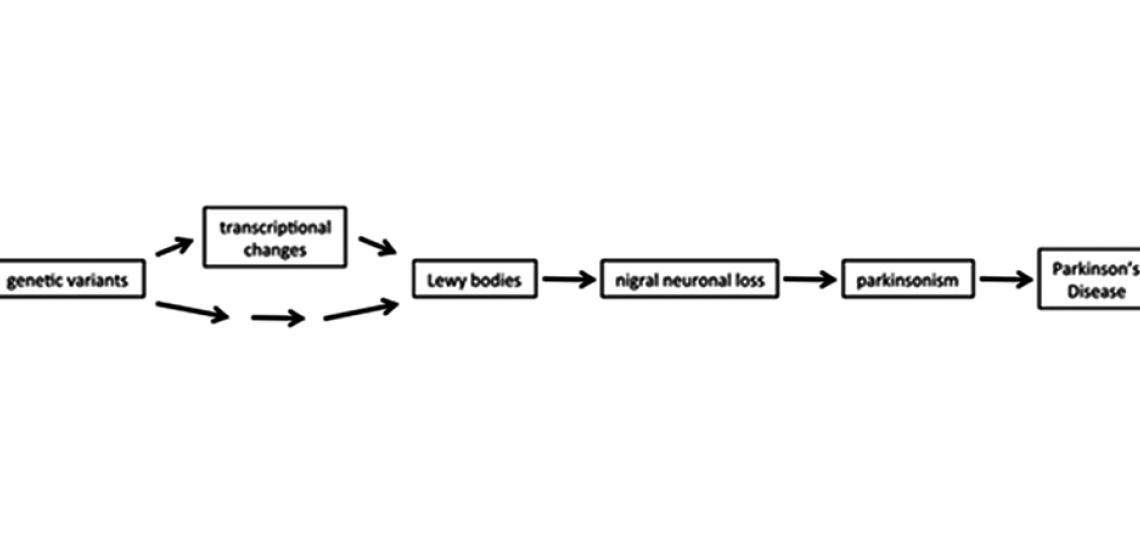
Exploring the contribution of common and rare genomic variation to disease susceptibility and progression.
We are also developing cohorts of well-characterized patients and families with PD and related movement disorders, including longitudinal data on disease progression, co-morbidities, and medication response. This work is performed in collaboration with clinician-investigators in the Baylor Parkinson's Disease Center and Movement Disorders Clinic and the University of Maryland Parkinson's Disease Center. Our investigation of these valuable study populations are complementary to the projects described above, allowing further exploration of the contribution of common and rare genomic variation to disease susceptibility and progression.








