About the Lab
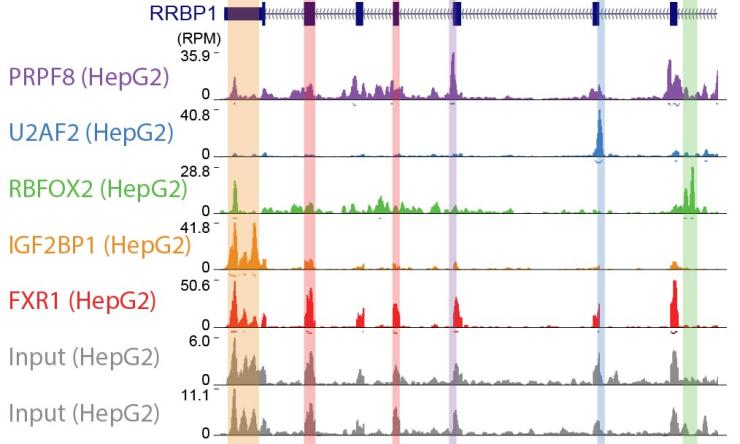
RNA binding proteins (RBPs) bind to RNA molecules in order to define the mature spliced sequence, direct nuclear export and sub-cellular localization, control translation initiation and efficiency, and regulate stability and ultimate decay. RNA processing plays critical roles in regulation of early development and differentiation of stem cells into mature lineages, and mis-regulation of RNA processing plays critical roles in cancer, neurodegenerative and other heritable diseases, and in both viral replication and anti-viral immune responses. Indeed, the emergence of genomics techniques have rapidly advanced our ability to identify genetic and transcriptomic causes of disease, and indeed there is now an ever-growing list of mutations in RBPs and RNA processing events causally linked to human diseases. However, it remains challenging to rapidly convert this genetic knowledge into the mechanistic understanding of the physiologically relevant mis-regulation required to develop therapeutic interventions, and to understand the complex regulatory networks controlled by the more than 1500 RBPs in the human genome. We utilize a mix of experimental and computational approaches to map RNA binding protein interaction networks and the regulatory roles of RBPs, in order to develop a global understanding of how RNA processing regulatory networks drive human physiology.
Research Projects
View a listing of current projects related to RNA binding proteins from the Van Nostrand Lab.

Publications
Our research projects and studies result in publications in PubMed and other scientific journals.
