Note:
The Macromolecular X-ray Crystallography Core and the Recombinant Protein Production and Characterization Core have merged into a new core, the “Recombinant Protein Production, Characterization, and Crystallography Core."
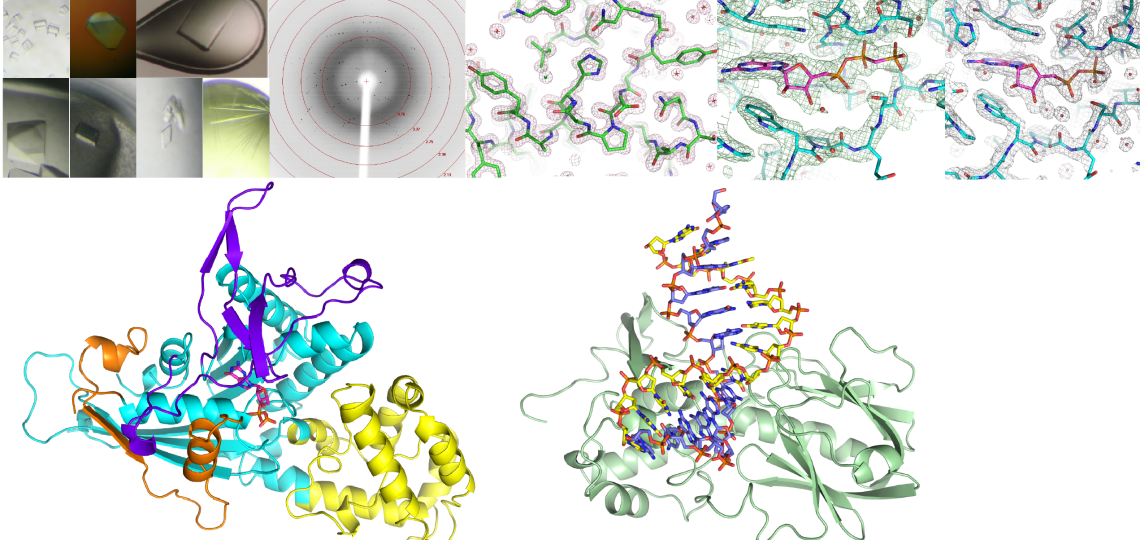
About the Core
Single crystal X-ray diffraction remains the most powerful technique to determine the three-dimensional structure of biologically important macromolecules and their functional ligand complexes at or near atomic resolution. The purpose of the Macromolecular X-ray Crystallography ATC is to provide a cost-efficient solution for scientists, researchers, and trainees at Baylor College of Medicine and the Texas Medical Center to pursue high-resolution structural studies. X-ray crystallography is not limited by the size or nature of the specimen, making it possible to determine the three-dimensional structure of nucleic acids and other small macromolecules to multi-subunit macromolecular assemblies.
X-ray Crystallography Advantages
One of the major advantages of X-ray crystallography is that atomic or near atomic resolution structural information can be obtained routinely, thereby revealing intimate details regarding the stereo-chemistry of the molecular interactions underlying the biological function of the macromolecule of interest. Finally, X-ray crystallography allows the three-dimensional structure determination of a macromolecule bound to an agonist or antagonist with little additional effort. The structure of such a complex is highly valuable, and the molecular insight thus obtained is frequently exploited for rationale structure-based drug design.
A new Formulatrix Rock Imager 2 capable of UV and Multi-fluorescence Imaging was acquired through an NIH Shared Instrumentation Grant Award (S10OD030246, Protein Crystallization Imager) and installed in 2022. Please remember to acknowledge our funding source in publications that use data produced by our core.
The BCM Macromolecular X-ray crystallography core is supported in part by an NIH Shared Instrumentation Grant Award (S10OD030246, Protein Crystallization Imager)
Macromolecular X-Ray Crystallography Core Protocols and Policies
Trace fluorescence labeling
Simple 0.1% protein labeling protocol
- Prepare 5 mM stock solution of a succinimidyl ester dye in DMSO
- Add appropriate amount of dye to protein solution for 0.1 % labeling of lysine residues assuming a 1:1 stoichiometric labeling efficiency to amine residues
- Wait 5 minutes for 90% dye binding
* There is no need for purification and samples are still fluorescent after 120 days
(Key Publications)
2022
Hu L, Salmen W, Chen R, Zhou Y, Neill F, Crowe JE Jr, Atmar RL, Estes MK, Prasad BVV. Atomic structure of the predominant GII.4 human norovirus capsid reveals novel stability and plasticity. Nat Commun. 2022 10;13(1):1241. PMID: 35273142; PMCID: PMC8913647.
Hu L, Salmen W, Sankaran B, Crawford SE, Estes MK, Prasad BVV (2022) Novel fold of rotavirus glycan-binding domain predicted by AlphaFold2 and determined by X-ray crystallography. Commun. Biol. 2022 5:419, PMID: 35513489; PMCID: PMC9072675
Mercado, J.M., Lee, S., Chang, C., Sung, N., Soong, L., Catic, A. and Tsai, F.T.F. Atomic Structure of the Leishmania spp. Hsp100 N-domain. Proteins 2022; 90:1242-1246. PMID: 35122310, PMCID: PMC9018533.
2021
Chamakuri, S., Lu, S., Ucisik, M.N., Bohren, K.M., Chen, Y.C., Du, H.C., Faver, J.C., Jimmidi, R., Li, F., Li, J.Y., Nyshadham, P., Palmer, S.S., Pollet, J., Qin, X., Ronca, S.E., Sankaran, B., Sharma, K.L., Tan, Z., Versteeg, L., Yu, Z., Matzuk, M.M.*, Palzkill, T.*, Young, D.W.* DNA-encoded chemistry technology yields expedient access to SARS-CoV-2 Mpro inhibitors. Proc. Natl. Acad. Sci. USA 2021 7;118(36) PMID: 34426525, PMCID: PMC8433497
Furey, I.M., Mehta, S.C., Sankaran, B., Hu, L., Prasad, B.V.V., and Palzkill, T. Local interactions with the Glu166 base and the conformation of an active site loop play key roles in carbapenem hydrolysis by KPC-2 β-lactamase. J. Biol. Chem. 2021 296:100799. PMID: 34022225, PMCID: PMC8189571.
Stojanoski, V., Hu, L., Sankaran, B., Wang, F., Tao, P., Prasad, B.V.V., and Palzkill, T. Mechanistic basis of OXA-48-like β-lactamases' hydrolysis of carbapenems. ACS Infect. Dis. 2021 7:445-460. PMID: 33492952, PMCID: PMC8571991.
Mehta, S.C., Furey, I.M., Pemberton, O.A., Boragine, D.M., Chen, Y., and Palzkill, T. KPC-2 β-lactamase enables carbapenem antibiotic resistance through fast deacylation of the covalent intermediate. J. Biol. Chem. 2021 296:100155. PMID: 33273017, PMCID: PMC7895804.
Taylor, D.M., Anglin, J., Hu, L., Wang, L., Sankaran, B., Wang, J., Matzuk, M.M., Prasad, B.V.V., Palzkill, T. Unique diacidic fragments inhibit the OXA-48 carbapenemase and enhance the killing of Escherichia coli producing OXA-48. ACS Infect. Dis. 2021 7:3345-3354. PMID: 34817169.
Maneix, L., Sweeney, M.A., Lee, S., Iakova, P., Moree, S.E., Sahin, E., Lulla, P., Yellapragada, S.V., Tsai, F.T.F. and Catic, A. The Mitochondrial Protease LonP1 Promotes Proteasome Inhibitor Resistance in Multiple Myeloma. Cancers 2021; 13:843. PMID: 33671345, PMCID: PMC7922145.
Yap, Z.Y., Efthymiou, S., Seiffert, S., Parra, K.V., Lee, S., Nasca, A., Maroofian, R., Schrauwen, I., Pendziwiat, M., Bhoj, E., Striano, P., Mankad, K., Vona, B., Cuddapah, S., Wagner, A., Alvi, J.R., Davoudi-Dehaghani, E., Fallah, M.-S., Gannavarapu, S., Lamperti, C., Legati, A., Murtaza, B.N., Rehman, M.U., Saeidi, K., Salpietro, V., Spiczak, S., Zeinali, S., Zeviani, M., SYNaPS Study Group, University of Washington Center for Mendelian Genomics, Helbig, I., Barakat, T.S., Ghezzi, D., Leal, S.M., Weber, Y., Houlden, H., Yoon, W.H. Bi-allelic Variants in OGDHL Cause a Neurodevelopmental Spectrum Disease Featuring Epilepsy, Hearing Loss, Visual Impairment, and Ataxia. Am. J. Hum. Genet. 2021; 108: 1-17. PMID: 34800363, PMCID: PMC8715183.
Lee, M.J., Bhattarai, D., Jang, H., Baek, A., Yeo, I.J., Lee, S., Miller, Z., Lee, S., Hong, J.T., Kim, D.-E., Lee, W., Kim, K.B. Macrocyclic Immunoproteasome Inhibitors as a Potential Therapy for Alzheimer’s Disease. J. Med. Chem. 2021; 64:10934-50. PMID: 34309393.
Yap, Z.Y., Park, Y. Wortmann, S.B., Gunning, A.C., Ezer, S., Lee, S., Duraine, L., Wilichowski, E., Wilson, K., Mayr, J.A., Wagner, M., Li, H., Kini, U., Black, E.D., Monaghan, K.G., Lupski, J.R., Ellard, S., Westphal, D.S., Harel, T., Yoon, W.H. Functional Interpretation of ATAD3A Variants in Neuro-mitochondrial Phenotypes. Genome Med. 2021; 13:55. PMID: 34845882, PMCID: PMC8042885.
Yap, Z.Y., Strucinska, K., Matsuzaki, S., Lee, S., Si, Y., Humphries, K., Tarnopolsky, M.A., and Yoon, W.H. A Biallelic Pathogenic Mutation in the OGDH Gene Results in a Neurological Disorder with Features of a Mitochondrial Disease. J. Inherit. Metab. Dis. 2021; 44:388-400. PMID: 32383294, PMCID: PMC7647956.
Rizzolo, K., Yu, A.Y.H., Ologbenla, A., Kim, S.-R, Zhu, H., Ishimori, K., Thibault, G., Leung, E., Zhang, Y., Teng, M., Haniszewski, M., Miah, N., Phanse, S., Minic, Z., Lee, S., Caballero, J.D., Babu, M., Tsai, F.T.F., Saio, T. and Houry, W.A. Functional Cooperativity Between the Trigger Factor Chaperone and the ClpXP Proteolytic Complex. Nature Commun. 2021; 12:281. PMID: 33436616, PMCID: PMC7804408.
Sharma, R., Choi, K.-J., Quan, M. D.,Sharma, S., Sankaran, B., Park, H., LaGrone, A., Kim, J. J., Mackenzie, K. R., Ferreon, A. C., Kim, C., and Ferreon, J. C. Liquid condensation of reprogramming factor KLF4 with DNA provides a mechanism for chromatin organization. Nature commun. 2021; 12, 5579, doi:10.1038/s41467-021-25761-7. PMID: 34552088, PMCID: PMC8458463.
2020
Soeung V, Lu S, Hu L, Judge A, Sankaran B, Prasad BVV, Palzkill T. A drug-resistant β-lactamase variant changes the conformation of its active-site proton shuttle to alter substrate specificity and inhibitor potency. J. Biol. Chem. (2020) 295:18239-55. PMID: 33109613.
Pan Y, Ren Z, Gao S, Shen J, Wang L, Xu Z, Yu Y, Bachina P, Zhang H, Fan X, Laganowsky A, Yan N, Zhou M. Structural basis of ion transport and inhibition in ferroportin. Nat. Commun. (2020) 11:5686. PMID: 33173040; PMCID: PMC7655804.
Shen J, Wu G, Tsai AL, Zhou M. Structure and mechanism of a unique diiron center in mammalian stearoyl-CoA desaturase. J. Mol. Biol. (2020) 432:5152-61. PMID:32470559; PMCID: PMC7483794.
Wang L, Qian H, Nian Y, Han Y, Ren Z, Zhang H, Hu L, Prasad BVV, Laganowsky A, Yan N, Zhou M. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature (2020) 581:329-32. PMID: 32433610; PMCID: PMC7255049.
Zhang H, Pan Y, Hu L, Hudson MA, Hofstetter KS, Xu Z, Rong M, Wang Z, Prasad BVV, Lockless SW, Chiu W, Zhou M. TrkA undergoes a tetramer-to-dimer conversion to open TrkH which enables changes in membrane potential. Nat. Commun. (2020) 11:547. PMID: 31992706; PMCID: PMC6987127.
Taylor DM, Anglin J, Park S, Ucisik MN, Faver JC, Simmons N, Jin Z, Palaniappan M, Nyshadham P, Li F, Campbell J, Hu L, Sankaran B, Prasad BVV, Huang H, Matzuk MM, Palzkill T. Identifying oxacillinase-48 carbapenemase inhibitors using DNA-encoded chemical libraries. ACS Infect. Dis. (2020) 6:1214-27. PMID: 32182432; PMCID: PMC7673237.
Brown CA, Hu L, Sun Z, Patel MP, Singh S, Porter JR, Sankaran B, Prasad BVV, Bowman GR, Palzkill T. Antagonism between substitutions in -lactamase explains a path not taken in the evolution of bacterial drug resistance. J. Biol. Chem. (2020) 295:7376-90. PMID: 32299911; PMCID: PMC7247304.
Wu F, Hua Y, Kaochar S, Nie S, Lin Y-L, Yao Y, Wu J, Wu X, Fu X, Schiff R, Davis CM, Robertson M, Ehli EA, Coarfa C, Mitsiades N, Song Y.* Discovery, structure-activity relationship and biological activity of histone-competitive inhibitors of histone acetyltransferases P300/CBP. J. Med. Chem. (2020) 63:4716-31. PMID: 32314924; PMCID: PMC7340344.
Bhattarai D, Lee MJ, Baek A, Yeo IJ, Miller Z, Baek YM, Lee S, Kim DE, Hong JT, Kim KB. LMP2 inhibitors as a potential treatment for Alzheimer’s disease. J. Med. Chem. (2020) 63:3763-83. PMID: 32189500.
Yu Z, Ku AF, Anglin JL, Sharma R, Ucisik MN, Faver JC, Li F, Nyshadham P, Simmons N, Sharma KL, Nagarajan S, Riehle K, Kaur G, Sankaran B, Srorl-Desmond M, Palmer SS, Young DW, Kim C, Matzuk MM. Discovery and characterization of bromodomain 2-specific inhibitors of BRDT. Proc. Natl. Acad. Sci. USA 2020 118(9) e2021102118 PMID: 33637650, PMCID: PMC7936271
2019
Tsai, J.T., Sung, N., Lee, J., Chang, C., Lee, S. and Tsai, F.T.F. Crystal Structure of the YcjX Stress Protein Reveals a Ras-Like GTP-Binding Protein. J. Mol. Biol. 2019, 431:3179-90."
2017
Lee J, Sung N, Yeo L, Chang C, Lee S, Tsai FT, Structural determinants for protein unfolding and translocation by the Hsp104 protein disaggregase, Bioscience Reports, 2017, 37(6):BSR20171399.
Lee J, Sung N, Mercado JM, Hryc CF, Chang C, Lee S, Tsai FT, Overlapping and Specific Functions of the Hsp104 N Domain Define Its Role in Protein Disaggregation, Scientific Reports 2017, 7(1):11184.
2016
Sung N, Lee J, Kim JH, Chang C, Tsai FT, Lee S, 2.4 Å resolution crystal structure of human TRAP1NM, the Hsp90 paralog in the mitochondrial matrix, Acta Crystallographica Section D 2016, 72(8):904-911.
Sung N, Lee J, Kim JH, Chang C, Joachimiak A, Lee S, Tsai FT, Mitochondrial Hsp90 is a ligand-activated molecular chaperone coupling ATP binding to dimer closure through a coiled-coil intermediate, Proceedings of the National Academy of Sciences of the United Stated of America 2016 113(11):2952-2957.